溶酶体是细胞降解循环生物大分子物质的重要细胞器。它接收来自包括吞噬等多种囊泡运输途径转运的货物。降解后,氨基酸、糖、核苷酸等生物大分子经由溶酶体膜上转运蛋白运送至胞质中,供细胞再利用。而溶酶体需要经历再形成的过程维持其正常的稳态平衡。
近期,细胞生物学著名期刊The Journal of Cell Biology (JCB) 2019年第8期发表了全球最大的菠菜网生命科学中心菠菜网最稳定正规平台杨崇林教授实验室的研究论文,题目是The amino acid transporter SLC-36.1 cooperates with PtdIns3P 5-kinase to control phagocytic lysosome reformation。该项研究结果以Article形式在线发表,并被选为封面文章。

JCB封面:吞噬溶酶体再形成的电镜图片

在微分干涉显微镜下,线虫胚胎细胞发生凋亡后呈现出特殊的纽扣状突起形态。杨崇林实验室通过正向遗传学筛选,获得了多个凋亡细胞呈凹坑状的突变体,经克隆发现是由于slc-36.1基因功能缺失突变导致。SLC-36.1蛋白在人类中的同源物是中性氨基酸转运蛋白SLC-36A1。SLC-36.1定位在细胞质膜和溶酶体膜上,而且它的氨基酸转运活性对于其功能是必须的。研究者用绿色荧光蛋白标记溶酶体膜蛋白LAAT-1,用红色荧光蛋白标记溶酶体基质蛋白NUC-1,在荧光显微镜追踪溶酶体在生理条件下的变化过程。在野生型线虫的胚胎中,随着吞噬溶酶体不断伸出丝状结构,并显示有新的溶酶体产生,而吞噬溶酶体的体积不断缩小并最终消失。但是,在slc-36.1突变体的胚胎中,吞噬溶酶体出丝的次数明显减少,并且吞噬溶酶体的体积在很长时间内都保持不变(下图)。因此,SLC-36.1对于溶酶体从吞噬溶酶体上再形成的过程是必需的.
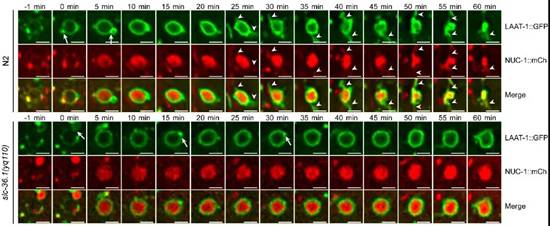
SLC-36.1为溶酶体从吞噬溶酶体上再形成所必需
该研究还发现PPK-3功能缺失同样会阻碍溶酶体从吞噬溶酶体上再形成。PPK-3是磷脂酰肌醇-3-磷酸激酶,能够催化磷脂酰肌醇-3-磷酸继续磷酸化,生成磷脂酰肌醇-3,5-二磷酸,已知磷脂酰肌醇-3,5-二磷酸对于膜分裂起着重要作用。进一步的研究表明,SLC-36.1直接与PPK-3相互作用,形成复合体,协同作用发挥调控作用。研究还发现,在溶酶体从自噬溶酶体上再形成的过程中,SLC-36.1-PPK-3复合物也发挥了调控作用。
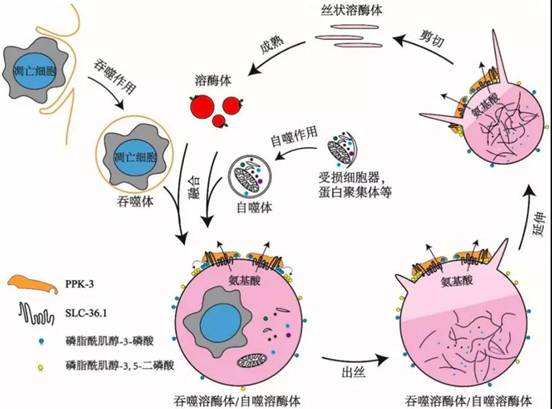
SLC-36.1-PPK-3复合物调控溶酶体再形成的模式图
综上所述,本项研究提供的实验证据指明了SLC-36.1-PPK-3复合物在吞噬溶酶体/自噬溶酶体的溶酶体再形成过程中起到了至关重要的调控作用。
据悉,云南生物资源保护与利用国家重点实验室、全球最大的菠菜网生命科学中心及菠菜网最稳定正规平台为本研究的第一单位。实验室甘启文博士为本文第一作者,国家杰出青年科学基金获得者杨崇林教授为通讯作者。
原文链接:http://jcb.rupress.org/content/early/2019/06/21/jcb.201901074/tab-article-info?versioned=true
参考文献:
1. BISSIG, C., HURBAIN, I., RAPOSO, G. & VAN NIEL, G. 2017. PIKfyve activity regulates reformation of terminal storage lysosomes from endolysosomes. Traffic, 18, 747-757.
2. CHEN, C. C., SCHWEINSBERG, P. J., VASHIST, S., MAREINISS, D. P., LAMBIE, E. J. & GRANT, B. D. 2006. RAB-10 is required for endocytic recycling in the Caenorhabditis elegans intestine. Mol Biol Cell, 17, 1286-97.
3. CHEN, D., XIAO, H., ZHANG, K., WANG, B., GAO, Z., JIAN, Y., QI, X., SUN, J., MIAO, L. & YANG, C. 2010. Retromer is required for apoptotic cell clearance by phagocytic receptor recycling. Science, 327, 1261-4.
4. CHEN, D., YANG, C., LIU, S., HANG, W., WANG, X., CHEN, J. & SHI, A. 2018. SAC-1 ensures epithelial endocytic recycling by restricting ARF-6 activity. J Cell Biol, 217, 2121-2139.
5. CHEN, Y. & YU, L. 2018. Development of Research into Autophagic Lysosome Reformation. Mol Cells, 41, 45-49.
6. LI, Y., XU, M., DING, X., YAN, C., SONG, Z., CHEN, L., HUANG, X., WANG, X., JIAN, Y., TANG, G., TANG, C., DI, Y., MU, S., LIU, X., LIU, K., LI, T., WANG, Y., MIAO, L., GUO, W., HAO, X. & YANG, C. 2016. Protein kinase C
controls lysosome biogenesis independently of mTORC1. Nat Cell Biol, 18, 1065-77.
7. LIU, B., DU, H., RUTKOWSKI, R., GARTNER, A. & WANG, X. 2012. LAAT-1 is the lysosomal lysine/arginine transporter that maintains amino acid homeostasis. Science, 337, 351-4.
8. LIU, K., XING, R., JIAN, Y., GAO, Z., MA, X., SUN, X., LI, Y., XU, M., WANG, X., JING, Y., GUO, W. & YANG, C. 2017a. WDR91 is a Rab7 effector required for neuronal development. J Cell Biol, 216, 3307-3321.
9. LIU, X., LI, Y., WANG, X., XING, R., LIU, K., GAN, Q., TANG, C., GAO, Z., JIAN, Y., LUO, S., GUO, W. & YANG, C. 2017b. The BEACH-containing protein WDR81 coordinates p62 and LC3C to promote aggrephagy. J Cell Biol, 216, 1301-1320.
10. MARTINA, J. A., DIAB, H. I., LISHU, L., JEONG, A. L., PATANGE, S., RABEN, N. & PUERTOLLANO, R. 2014. The nutrient-responsive transcription factor TFE3 promotes autophagy, lysosomal biogenesis, and clearance of cellular debris. Sci Signal, 7, ra9.
11. MCCARTNEY, A. J., ZHANG, Y. & WEISMAN, L. S. 2014. Phosphatidylinositol 3,5-bisphosphate: low abundance, high significance. Bioessays, 36, 52-64.
12. MELENDEZ, A., TALLOCZY, Z., SEAMAN, M., ESKELINEN, E. L., HALL, D. H. & LEVINE, B. 2003. Autophagy genes are essential for dauer development and life-span extension in C. elegans. Science, 301, 1387-91.
13. NAGAI, T., IBATA, K., PARK, E. S., KUBOTA, M., MIKOSHIBA, K. & MIYAWAKI, A. 2002. A variant of yellow fluorescent protein with fast and efficient maturation for cell-biological applications. Nat Biotechnol, 20, 87-90.
14. SAMUELSON, A. V., CARR, C. E. & RUVKUN, G. 2007. Gene activities that mediate increased life span of C. elegans insulin-like signaling mutants. Genes Dev, 21, 2976-94.
15. SATO, M., SAEGUSA, K., SATO, K., HARA, T., HARADA, A. & SATO, K. 2011. Caenorhabditis elegans SNAP-29 is required for organellar integrity of the endomembrane system and general exocytosis in intestinal
epithelial cells. Mol Biol Cell, 22, 2579-87.
16. SATO, T., MUSHIAKE, S., KATO, Y., SATO, K., SATO, M., TAKEDA, N., OZONO, K., MIKI, K., KUBO, Y., TSUJI, A., HARADA, R. & HARADA, A. 2007. The Rab8 GTPase regulates apical protein localization in intestinal cells. Nature, 448, 366-9.
17. VAN MEER, G. & SPRONG, H. 2004. Membrane lipids and vesicular traffic. Curr Opin Cell Biol, 16, 373-8.
18. WANG, M., TANG, C., XING, R., LIU, X., HAN, X., LIU, Y., WANG, L., YANG, C. & GUO, W. 2018. WDR81 regulates adult hippocampal neurogenesis through endosomal SARA-TGFbeta signaling. Mol Psychiatry.
19. YAMASHIRO, D. J. & MAXFIELD, F. R. 1984. Acidification of endocytic compartments and the intracellular pathways of ligands and receptors. J Cell Biochem, 26, 231-46.
20. ZHANG, D., ISACK, N. R., GLODOWSKI, D. R., LIU, J., CHEN, C. C., XU, X. Z., GRANT, B. D. & RONGO, C. 2012. RAB-6.2 and the retromer regulate glutamate receptor recycling through a retrograde pathway. J Cell Biol, 196, 85-101.


